VCPA
VCPA - SNP/Indel Variant Calling Pipeline and data management tool used for analysis of whole genome and exome sequencing (WGS/WES) for the Alzheimer’s Disease Sequencing Project

VCPA consists of two independent but linkable components: pipeline and tracking database. The pipeline is coded in Workflow Description Language (https://bitbucket.org/NIAGADS/vcpa-pipeline/src/master/) and is fully optimized for the Amazon elastic compute cloud environment (ami-acc840d3). Documentation on how to run VCPA is availalble here (https://bitbucket.org/NIAGADS/vcpa-pipeline/src/master/doc/).
VCPA includes steps for processing raw sequence reads including read alignment, and all the way up to variant calling using GATK. The tracking database allows users to dynamically view the statuses of jobs running and the quality metrics reported by the pipeline. Users can thus monitor the production process and diagnose if any problem arises during the procedure. All quality metrics (>100 collected per processed genome) are stored in the database, thus facilitating users to compare, share and visualize the results. To summarize, VCPA is functionally equivalent to the CCDG/TOPMed pipeline. Together with the dockerized database (also available as Amazon Machine Image), users can easily process any WGS/WES data on Amazon cloud with minimal installation.
VCPA is released under the MIT license and is available for academic and nonprofit use for free.
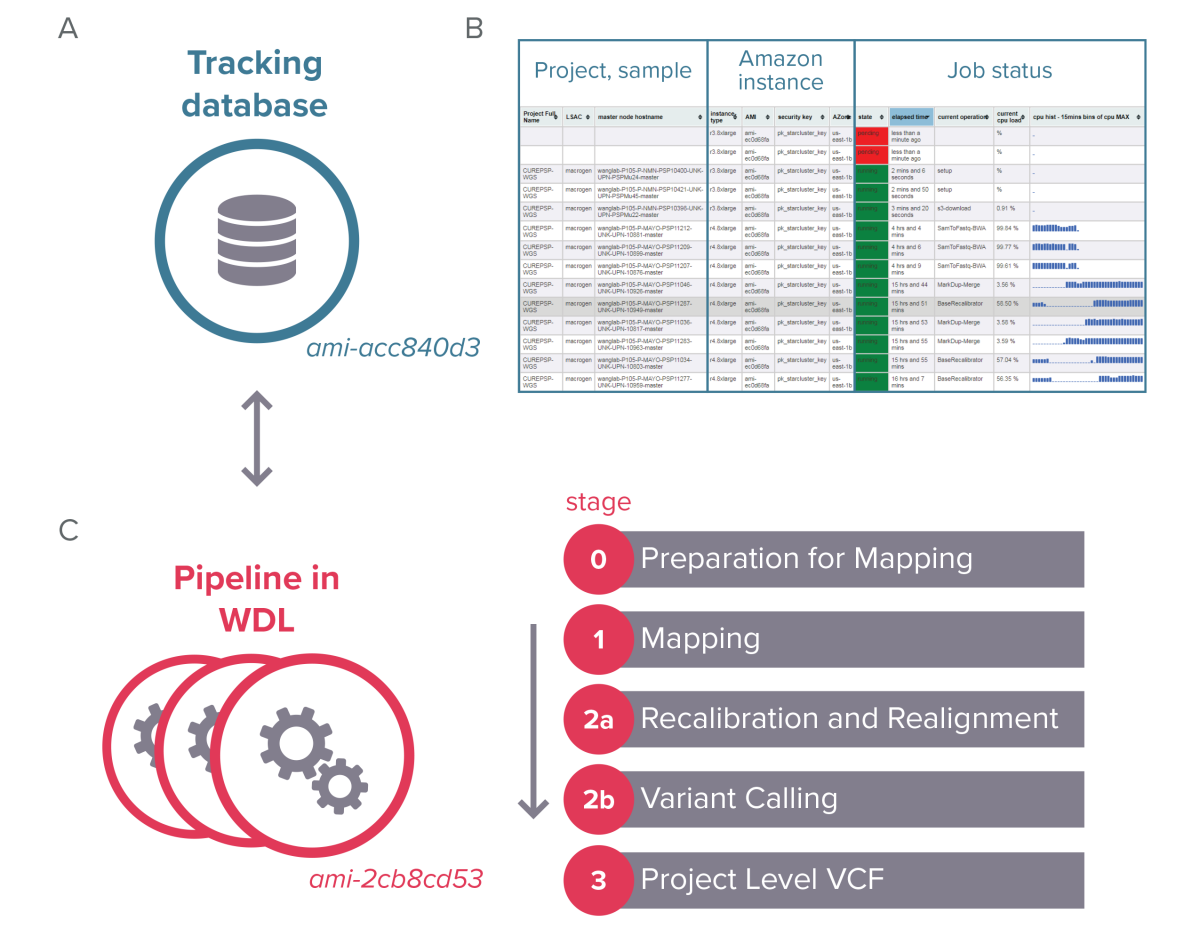
Figure 1: A) VCPA architecture; B) Dynamic view of job status; C) Pipeline overview.
To cite VCPA, please use this citation:
Yuk Yee Leung, Otto Valladares, Yi-Fan Chou, Han-Jen Lin, Amanda B Kuzma, Laura Cantwell, Liming Qu, Prabhakaran Gangadharan, William J Salerno, Gerard D Schellenberg, Li-San Wang. VCPA: genomic variant calling pipeline and data management tool for Alzheimer’s Disease Sequencing Project, Bioinformatics, 2018 https://doi.org/10.1093/bioinformatics/bty894
